This note is a high-fidelity Markdown migration of the Maximum Likelihood and Machine Learning chapter from the LaTeX source.
Parent map: index Prerequisites: probability-and-mathstats, linear-regression, causal-inference
Concept map
flowchart TD A[Maximum Likelihood] --> B[Score / Fisher Information] A --> C[QMLE / Robust SEs] A --> D[Binary / Discrete Choice] D --> E[Ordered / Unordered Models] A --> F[Counts / Rates / Truncation] A --> G[GLM Theory] H[Machine Learning] --> I[Supervised Learning] I --> J[Regularization] I --> K[Classification Metrics] H --> L[Neural Networks] H --> M[Unsupervised Learning]
Maximum Likelihood
Setup
Let be a sequence of iid rv’s with common CDF . We want to estimate .
Since the rv’s are IID, we can write
L:
We usually work with the log of this , and drop the conditioning on though strictly speaking
is the estimator that maximises the conditional log-likelihood estimator.
solves the first-order conditions
conditional density:
Log likelihood:
Maximising this w.r.t. yields , .
gradient vector
Evaluated at , this is the efficient score.
local maximum that solves the FOCs , IOW
Variance estimate ;
Estimated as
For OLS, parameter vector .
Scores:
Information Matrix / Variance
Properties of Maximum Likelihood Estimators
, probability of missing the true parameter goes to zero.
Equivalently,
conditions: (1) , (2) is twice-differentiable Variance of MLE is the Cramer-Rao Bound; the asymptotic variance of the MLE is at least as small as that of any other consistent estimator. Let the pdf of the r.v. be for some . Let be an unbiased estimator for . Suppose the derivative can be passed under the integral and and suppose the fisher information
is finite. Then,
If are iid with common density , the implied bound on the variance is Let be iid random variables with common density . A sequence of estimates , a function that satisfies
whatever the true value of is, is said to be asymptotically efficient. Suppose are iid with density where is an unknown function. Next, we pretend to know the infinite dimensional parameter up to a finite dimensional parameter , in which we have a fully parametric finite-dimensional parameter , thus we can calculate the Cramer-Rao Bound.
Partitioning the information matrix for and its inverse in
The Cramer-Rao bound implies that
This is true for any parametrisation of the unknown function . The lowest possible variance for any estimator for that does not use knowledge of has to be at least as high as the lowest variance we can get if we know more, that is, the Cramer-Rao bound for any parametric submodel. So, the semiparametric efficiency bound is the largest lower-bound we can get for any parametric submodel.
Suppose we have a candidate Estimator and a given parametrisation . Then,
For any estimator we can calculate the left hand side, for any parametrization we can calculate the right hand side, so if we find an estimator and a parametrization that the two are equal we have found the efficiency bound. Let where is bijective, continuous, and differentiable. Let be the MLE of . Then, is the MLE of . For a regression of the form , one can estimate multiple ‘marginal effects’. For the special case where , , but this is not generically true.
Average Marginal Effect (AME):
Marginal Effect at Mean (MEM) Suppose is a sufficient statistic for . Then,
depends on the data only through , the sufficient statistic.
QMLE / Misspecification / Information Theory
If model is misspecified,
The MLE converges to the best fitting for the population (pseudo-true value)
For the linear exponential family, the quasi-MLE is consistent even when the density is partially misspecified.
Robust Standard Errors
Asymptotic distribution of QMLE
where
let be the assumed joint density, and let be the true density. Then,
Minimised when . QMLE minimises distance between and . Notation denotes ‘information lost when is used to approximate ‘.
Discrete version illustrates links to Entropy
and with equality IFF .
‘distance’, unlike Euclidian Distance, is not the same between as ; i.e. it is directional. Akaike showed that using K-L model selection entails finding a good estimator for
where are independent, random samples from the same distribution and expectations are taken w.r.t. the true distribution . Estimating this quantity for each model is biased upwards. An approximately unbiased estimator of the above target quantity is
For a general class of maximum-likelihood models,
For linear regression models, this simplifies to
Testing
To test the hypothesis against the alternative, there are three classical tests.
We partition the parameter -vector into two parts s.t. the dimensions of the two sub-vectors s.t. . is a nuisance parameter: its value is not restricted under the null.
Let be the unrestricted MLEs. If we are testing the restriction , then the restricted parameter vector is . IOW, test .
If null is true, at restricted model () should not be much smaller than at the unrestricted model (.
Under the null, (where is the number of restrictions being tested). If the limiting is maximised at , the derivative of the wrt at that point should be close to zero.
Under the null, (where is the number of restrictions being tested). Unrestricted estimates of should be close to zero.
Where is the top-left of the information matrix (corresponding with the restricted parameters). Under the null, .
alternatively,
where are restrictions,
evaluated at . McFadden’s Pseudo-
Binary Choice
Estimate the probability using OLS . , so heteroskedasticity is mechanically present unless all coefficients are zero.
Logistic regression fits , where is the link function that scales onto the probability scale. Alternatively, one can use . For , assume latent index model $Y_i^* = X_i’\beta
- \epsilon_iY_i \defeq \Indic{Y_i^* > 0}Y_i\mathcal{L}=\prod_{i=1}^{N} \pi_{i}^{Y_{i}}\left(1-\pi_{i}\right)^{1-Y_{i}}$.
Symmetric CDFs. Let
Probit:
Logit:
=
| Model | Marginal Effect | |
|---|---|---|
| Logit | ||
| Probit | ||
| Clog-log | ||
| LPM |
Let be the density and CDF evaluated at .
Sample Score solves
Variance:
Marginal effect:
Since for the logistic CDF, , the score and hessian can be written as
Discrete Choice
In many Additive Random Utility Models (ARUMs), is logistic for multivariate extensions to logit. This assumes that the errors themselves are distributed Gumbel/ type 1 extreme-value distribution
and .
Ordered
Random utility with multiple cutoffs , where .
Define latent variable, and
which means
specifications yield a likelihood that is simply the product of binary logit/probit models that switch between adjacent categories for each observation.
Marginal effects are of the form
Unordered
Multinomial distribution
where we adopt normalisation for identification.
Coefficient interpretation:
which implies that the log-odds ratio is linear in . Permits incorporation of choice-varying predictors , nests MNL.
Log likelihood of the form
Relative risk independent of other choices ; choices are series of pairwise comparisons. . IoW .
where
Counts and Rates
Counts
Poisson specification: . Yields log density
Score:
solves
Hessian
Assumes .
Marginal Effect: Since for poisson, . Parameters can be interpreted as semi elasticities, since
define a bernoulli for observation, and specify separate models for zero and nonzero data, with potentially different covariates on and . Yields the following (difficult to maximise) likelihood
. Let , ,
Rates
Survival:
Hazard: ;
Cumulative Hazard .
Proportional Hazard Models
Conditional hazard rate can be factored as
baseline hazard ( for exponential and for Weibull).
| Parametric model | Hazard | Survival |
|---|---|---|
| Exponential | ||
| Weibull | ||
| Generalised Weibull | ||
| Gompertz |
For survival models with censoring, Likelihood is often written as
where is a right-censoring indicator and is the observed time.
Weibull Density: . . Specify , so . Then, the log-likelihood is
FOCs are
Model needs to be correctly specified to be consistent. Unlike OLS or poisson.
Truncation and Censored Regressions
If a c.r.v and is truncated at ,
For the truncated normal distribution where is truncated at ,
,
where and is the inverse Mills ratio / Hazard function.
Tobit Regression
Censored s.t. and (i.e. is censored from below at c).
Truncated MLE maximises
with and denoting the density and distribution of respectively.
Type-I Tobit assumes is normally distributed, which gives us the following likelihood
Censored Regression
Consider a model and is not observed if .
Yields log-likelihood
Generalised Linear Models Theory
Semi-robust likelihoods belong to the Linear exponential Family of the following form:
Response observations are realisations of random variables with densities of the form
is called the canonical / natural parameter, is the dispersion parameter. ,
Linear predictor specifying the variation in accounted for by known covariates. is a transformation of the mean that addresses scaling. It is so called because it links the expected value of the response variable to the explanatory covariates.
Since , under a canonical link ( ), .
ML estimation
log likelihood
Score function
The FoC can be written as
where is a weight matrix (which depends on ). The fitted value
By a first-order taylor expansion, define
This gives us an update rule
repeat until convergence .
Model Density Link
OLS Gaussian Identity Logistic Binomial Logistic Logistic Binomial Normal Poisson Poisson Log
Machine Learning
Supervised Learning
Every Supervised ML algorithm essentially involves a function class and a regulariser that expresses the complexity of the representation. Then, two steps
conditional on a level of complexity, choose best in-sample loss-minimising function
Estimate the ‘optimal’ level of complexity using empirical tuning
| Aspect | Discriminative model | Generative model |
|---|---|---|
| Goal | Directly estimate | Estimate to deduce |
| What is learned | Decision boundary | Probability distribution of the data |
| Examples | Regressions, SVM | GDA, Naive Bayes |
that takes as inputs predicted value and real data value and outputs how different they are.
Least Squares:
Logistic:
Hinge:
Cross Entropy:
| Predictor class | Examples | Regulariser / tuning parameters |
|---|---|---|
| Global / parametric predictors | Linear and generalisations | Subset selection ; LASSO ; Ridge ; Elastic Net |
| Local / nonparametric predictors | Decision / Regression trees; Random forest; Nearest neighbours; Kernel regression | Tree depth, number of nodes/leaves, minimal leaf size, information gain at splits; number of trees; number of variables used in each tree; bootstrap sample size; number of neighbours; kernel bandwidth |
| Mixed predictors | Neural networks (including deep and convolutional); splines | Number of layers, number of neurons per layer, connectivity between neurons; number of knots and order |
| Combining predictors | Bagging; Boosting; Ensemble methods | Number of draws, bootstrap size, individual tuning parameters; learning rate and number of iterations; ensemble weights |
Reference: Mullainathan and Spiess (2017), Table 2.
take a unit hypercube in dimension and we put another hypercube within it that captures a fraction of observations within the cube. Each edge will be . For moderately high dimensions , . Need 80% data to cover 10% of sample.
Define d(p,N) as distance from the origin to the closest point. (closest point closer to the boundary than to the origin).
Regularised Regression
In general, we want to impose a penalty for model complexity in order to minimise MSE (trade off some bias for lower variance).
Estimate the following regression
where is a standardized design matrix (s.t. all Xs have unit variance). Let be the SVD of .
Then, ridge coefficients can also be written as
This can be used to compute the ridge coefficient efficiently for a fine grid of s.
Compute SVD of and save Compute and store for For each compute compute
The solution vector is ‘biased’ towards the leading right singular vectors of , which gives it the property of a ‘smoothed’ Principal Components regression. For Ridge regression,
Where s are the eigenvalues of the Covariance Matrix.
More generally, for any smoother matrix , , which may not be an integer for semi/non-parametric smoothers. In the special parametric case of OLS, , so the DoF is simply . Consider the objective function
fit using sequential coordinate descent. Coefficient vector is soft-thresholded:
where is a standardized design matrix (s.t. all Xs have unit variance), and is the norm. both cases, pick tuning parameter using cross-validation.
ML analogue to LASSO. Define
where
Combines ridge regression and the lasso by adding a penalty to the LASSO’s objective function
Generalise the penalty to a class of penalty functions of the form
where is a diagonal matrix whose diagonal elements are functions of the squared singular values.
Define the following objective function
Where is a diagonal matrix with diagonal entries equalling . This penalty term gives no weight to the component of that aligns with the first right singular vector of (i.e. the first principal component. This gives it better predictive accuracy in some settings.
Comparing principal-coordinate predictions of ridge and pcLASSO:
The latter corresponds to a more aggressive form of shrinkage towards the leading singular vectors.
Classification
Training sample where (can relabel to Bernoulli). A predictor , where the labels are produced by an (unknown) classifier . Let be an (unknown) distribution on . The error of w.r.t. is defined by
The empirical risk is defined as
A perfect classifier (in the sense that does not exist, so we aim for Probably Approximately Correct (PAC) learners that have w.p. . The space of models is restricted to be in finite set . It can be shown that , if , then where
loosely represents the expressive capacity of a set of functions.
Consider points and the set
we say that m shatters all the points if , i.e. all combinations are possible.
Linear functions can shatter 2 points.
The VC dimension of is
Let . A linear classifier can then be written as where
Suppose a hyperplane s.t. .
The hyperplane that separates the data and maximises the ‘margin’ is given by minimising . Typically, the function space is large and complex, so a natural idea is to learn iteratively. Loosely, estimate a model for from , which produces error . Next, estimate for from , which produces , and so on. So, after steps,
where the first error is and so on, and can also be seen as the gradient associated with the quadratic loss function, . So, an equivalent representation is
where is a space of ‘weak learners’ (typically step functions). To ensure ‘slow’ learning, one typically applies a shrinkage parameter . Arthur Charpentier’s series on probabilistic foundations of econometrics and machine learning covers this topic and includes a useful bibliography.
Goodness of Fit for Classification
calibration: Bin predicted probabilities into bins , and within each compute (average predicted probability) and . Plot the two average against each other. In a well calibrated model, the binned averages trace the identity line.
discrimination: Discrimination is a measure of whether observations have high , and correspondingly values have low . Many measures; listed below Observed
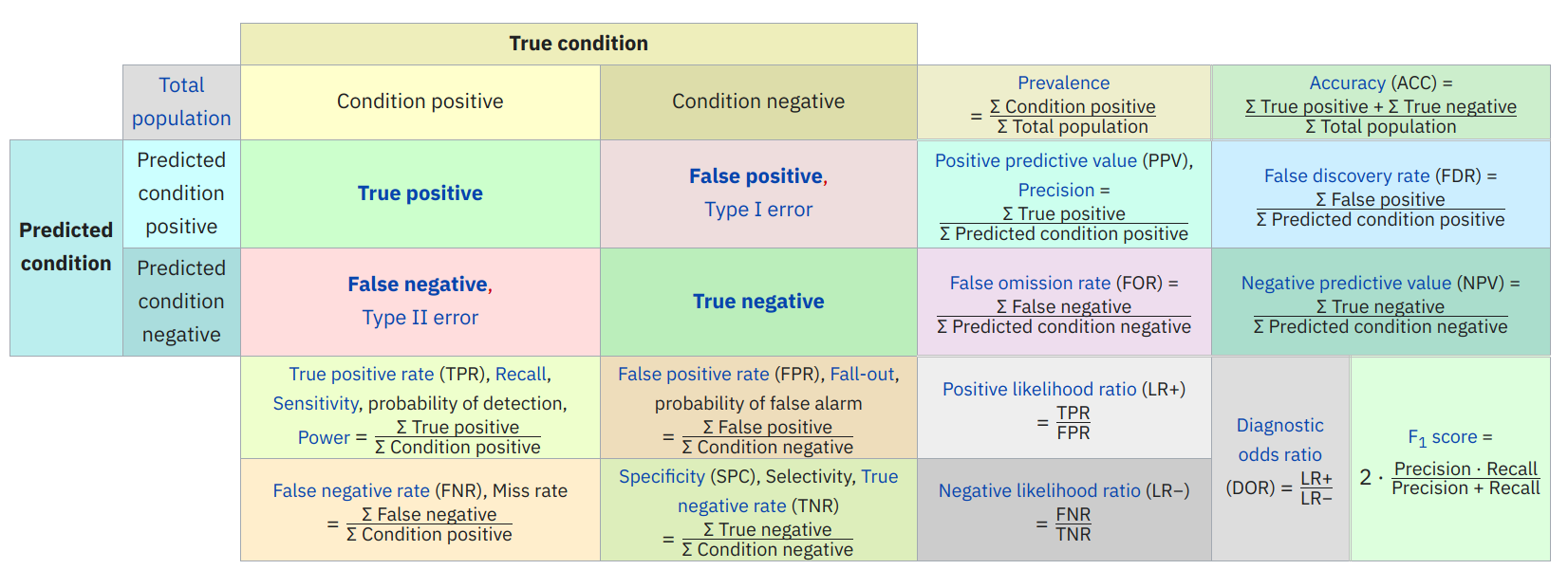
Predicted positive () True Positive (TP) False Positive (FP) Predicted negative () False Negative (FN) True Negative (TN) Total Positive(P) Total Negative(N) Accuracy = - Overall performance
Precision = - How accurate positive predictions are
Sensitivity = Recall = True positive Rate = - Coverage of actual positive sample
Specificity = True Negative Rate = - Coverage of actual negative sample
Brier Score =
F1 Score = : hybrid metric for unbalanced classes
Is the plot of TPR vs FPR by varying the threshold .
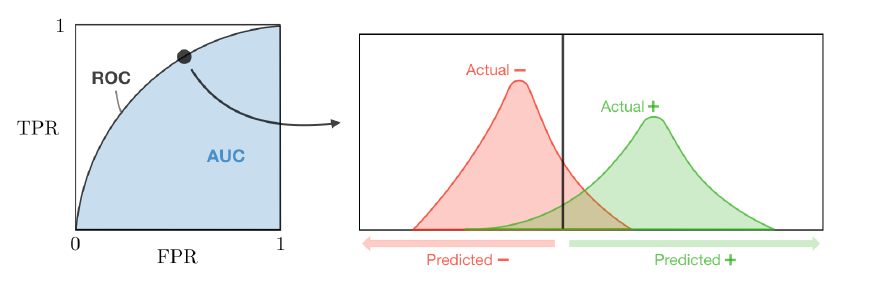
Suppose we have a training set , a test point , and a tree predictor
Equivalently,
where is partitioned into leaves , where leaves are constructed to maximise heterogeneity between nodes . Do this until all leaves have minimum leaf size observations. Regression trees overfit, so we need to use cross-validation + other tricks.
Random forests build and average many different trees by
Bagging / subsampling training set (Breiman)
Selecting the splitting variable at each step from out of randomly drawn features (Amit and Geman)
Generalised nonparametric regression with many ‘layers’, with components outline in the referenced figure. For the layer of the network and hidden layer of the unit, we have
where are the weight (coefficient), bias (intercept) and output respectively.
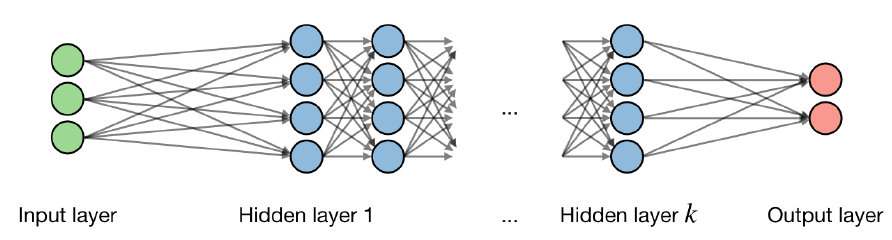 Activation functions are used at the end of a hidden layer to introduce non-linearities into the model. Common ones are
Activation functions are used at the end of a hidden layer to introduce non-linearities into the model. Common ones are
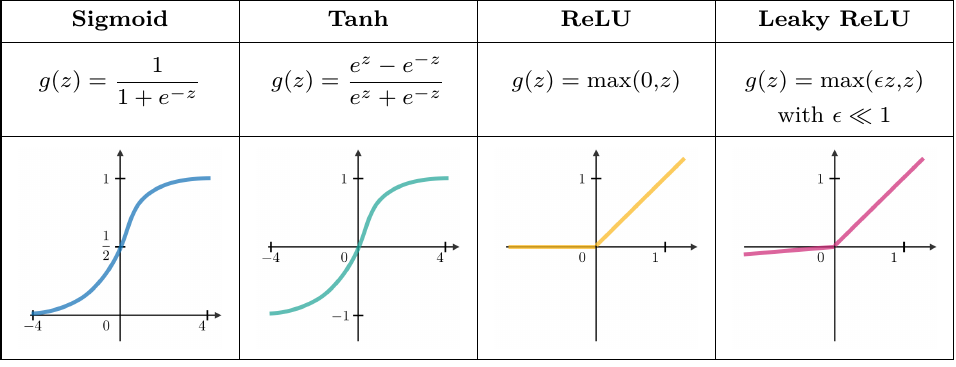
Networks frequently use the cross-entropy loss function.
Learning rate is denoted by , which is the pace at which the weights get updated. This can be fixed or adaptively changed using ADAM.
Back-propagation is a method to update the weights in the neural net by taking into account the actual output and desired output. The derivative with respect to weight is computed using the chain rule and is of the following form
So the weight is updated
- Take a batch of training data.
- Perform forward propagation to compute corresponding loss.
- Perform back propagation to compute gradients.
- Use the gradients to update the weights over the network.
Unsupervised Learning
There is no distinction between a label/outcome and predictor in a wide variety of problems. The goal of unsupervised methods is to characterise the joint distribution of the data using latent factors, clusters, etc.
Original data in . We approximate orthogonal unit vectors and associated scores ( weights ) to minimise reconstruction error
where subject to the constraint that the smoother matrix is orthonormal. Equivalently, the objective function can be written as
where is with in its rows.
The optimal solution sets each to be the l-th eigenvector of the empirical covariance matrix. Equivalently, , which contains the eigenvectors with the largest eigenvalues of empirical covariance matrix . If we rank singular values of the data matrix , we can construct a rank approximation, the truncated SVD
This is identical to the optimal reconstruction .
The error is the sum of remaining eigenvalues of the covariance matrix. Total variance explained = (sum of included eigenvalues)/(sum of all eigenvalues)